Why the long face?
Syngnathid fish – that is seahorses, pipefish, and seadragons – have many spectacularly derived morphological and behavioral adaptations. Most well-known is probably the sex-role reversal and male pregnancy, with females laying their eggs in a brood pouch situated on the tail or the trunk of the male (depending on the species), where the embryos develop into juvenile fish.
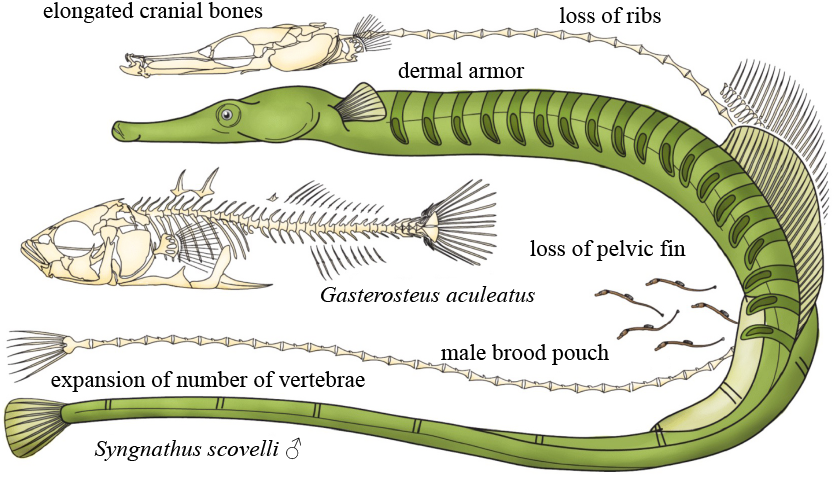
Syngnathid-specific adaptations, with comparison of a Gulf pipefish (Syngnathus scovelli) and a three-spine stickleback (Gasterosteus aculeatus). Modified figure from the Cresko Lab paper on the Gulf pipefish genome: Small et al. (2016) Genome Biology (https://doi.org/10.1186/s13059-016-1126-6) under CC BY 4.0.
Other striking adaptations include the expansion of number of vertebrae, the loss of all ribs, the evolution of body armor, the loss pelvic fins, and the main target for my current studies: the snout-like mouth. The snout is not really the mouth per se, since the jaws are situated at the end of it, organized very differently from other fish, with the lower jaw at a sharp angle upwards. This, together with the long snout, allow for very efficient “pivot suction feeding”, so that the syngnathids can rapidly open their mouth and create underpressure that sucks in their prey.
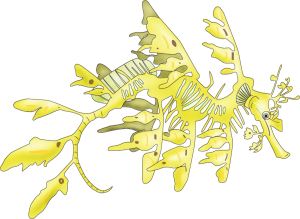
A leafy seadragon (Phycodurus eques), spectacularly camouflaged as floating seaweed. Illustration (c): Allison Fuiten.
But how does this strange craniofacial morphology develop? When – and where in the evolutionary tree – did it evolve? Developmental pathways controlling craniofacial development are incredibly conserved across all vertebrates, so the same genes that shape your mouth are involved in shaping the beaks of birds, and the snouts of pipefish.
I am therefore mainly focusing on investigating whether new ways of regulating the genes in these developmental pathways have changed. Could the syngnathid fish have lost some key elements to regulate gene expression, or have they gained new, lineage-specific ones?

The trunk-brooding bluestripe pipefish (Doryrhamphus excisus). Illustration (c): Allison Fuiten.
Comparative genomics
One way of assessing lineage-specific losses and gains of regulatory elements is by comparing regions of conserved DNA sequence across the genomes of syngnathid fish and other vertebrates. For this, I am aligning the whole genomes of tens of bony fish species, coelacanth, and representatives of tetrapod lineages such as frogs, snakes, birds, and mammals.
Experimental validation with CRISPR/Cas9

The zebrafish was first introduced as a model system at the University of Oregon. This picture also happens to be the fruit of my very first attempt at scientific illustrations, made during a workshop with Allison Fuiten.
I will also experimentally validate candidate regulatory elements using CRISPR/Cas9. In case of syngnathid-specfic losses of regulatory elements, I will test the phenotypic effects by knocking those elements out in three-spine stickleback Gasterosteus aculeatus and zebrafish Danio rerio.
